-
This image depicts a Petri dish containing a buffered charcoal yeast extract (BCYE) agar medium, which had been inoculated with Gram-negative Francisella tularensis live vaccine strain (LVS) bacteria. F. tularensis is the pathogen responsible for causing the disease tularemia. This was the appearance of the colonial growth after 72 hours of incubation. What is tularemia?Tularemia, also known as rabbit fever, is a disease caused by the bacterium Francisella tularensis. Tularemia is typically found in animals, especially rodents, rabbits, and hares. Tularemia is usually a rural disease and has been reported in all U.S. states except Hawaii.How do people become infected with tularemia?Typically, people become infected through the bite of infected insects (most commonly, ticks and deerflies), by handling infected sick or dead animals, by eating or drinking contaminated food or water, or by inhaling airborne bacteria.Created: 2009
-
This image depicts a Petri dish containing a chocolate agar medium, which had been inoculated with Gram-negative Francisella tularensis live vaccine strain (LVS) bacteria. F. tularensis is the pathogen responsible for causing the disease tularemia. This was the appearance of the colonial growth after 72 hours of incubation. What is tularemia?Tularemia, also known as rabbit fever, is a disease caused by the bacterium Francisella tularensis. Tularemia is typically found in animals, especially rodents, rabbits, and hares. Tularemia is usually a rural disease and has been reported in all U.S. states except Hawaii.How do people become infected with tularemia?Typically, people become infected through the bite of infected insects (most commonly, ticks and deerflies), by handling infected sick or dead animals, by eating or drinking contaminated food or water, or by inhaling airborne bacteria.Created: 2009
-
This image depicts a Petri dish containing a sheeps blood agar (SBA) medium, which had been inoculated with Gram-negative Francisella tularensis live vaccine strain (LVS) bacteria. F. tularensis is the pathogen responsible for causing the disease tularemia. This was the appearance of the colonial growth after 48 hours of incubation. What is tularemia?Tularemia, also known as rabbit fever, is a disease caused by the bacterium Francisella tularensis. Tularemia is typically found in animals, especially rodents, rabbits, and hares. Tularemia is usually a rural disease and has been reported in all U.S. states except Hawaii.How do people become infected with tularemia?Typically, people become infected through the bite of infected insects (most commonly, ticks and deerflies), by handling infected sick or dead animals, by eating or drinking contaminated food or water, or by inhaling airborne bacteria.Created: 2009
-
This image depicts a Petri dish containing a chocolate agar medium, which had been inoculated with Gram-negative Francisella tularensis live vaccine strain (LVS) bacteria. F. tularensis is the pathogen responsible for causing the disease tularemia. This was the appearance of the colonial growth after 48 hours of incubation. What is tularemia?Tularemia, also known as rabbit fever, is a disease caused by the bacterium Francisella tularensis. Tularemia is typically found in animals, especially rodents, rabbits, and hares. Tularemia is usually a rural disease and has been reported in all U.S. states except Hawaii.How do people become infected with tularemia?Typically, people become infected through the bite of infected insects (most commonly, ticks and deerflies), by handling infected sick or dead animals, by eating or drinking contaminated food or water, or by inhaling airborne bacteria.Created: 2009
-
This image depicts a Petri dish containing a buffered charcoal yeast extract (BCYE) agar medium, which had been inoculated with Gram-negative Francisella tularensis live vaccine strain (LVS) bacteria. F. tularensis is the pathogen responsible for causing the disease tularemia. This was the appearance of the colonial growth after 48 hours of incubation. What is tularemia?Tularemia, also known as rabbit fever, is a disease caused by the bacterium Francisella tularensis. Tularemia is typically found in animals, especially rodents, rabbits, and hares. Tularemia is usually a rural disease and has been reported in all U.S. states except Hawaii.How do people become infected with tularemia?Typically, people become infected through the bite of infected insects (most commonly, ticks and deerflies), by handling infected sick or dead animals, by eating or drinking contaminated food or water, or by inhaling airborne bacteria.Created: 2009
-
This image depicts a Petri dish containing a chocolate agar medium, which had been inoculated with Gram-negative Francisella tularensis live vaccine strain (LVS) bacteria. F. tularensis is the pathogen responsible for causing the disease tularemia. This was the appearance of the colonial growth after 24 hours of incubation. What is tularemia?Tularemia, also known as rabbit fever, is a disease caused by the bacterium Francisella tularensis. Tularemia is typically found in animals, especially rodents, rabbits, and hares. Tularemia is usually a rural disease and has been reported in all U.S. states except Hawaii.How do people become infected with tularemia?Typically, people become infected through the bite of infected insects (most commonly, ticks and deerflies), by handling infected sick or dead animals, by eating or drinking contaminated food or water, or by inhaling airborne bacteria.Created: 2009
-
This image depicts a Petri dish containing a sheeps blood agar (SBA) medium, which had been inoculated with Gram-negative Francisella tularensis live vaccine strain (LVS) bacteria. F. tularensis is the pathogen responsible for causing the disease tularemia. This was the appearance of the colonial growth after 24 hours of incubation. What is tularemia?Tularemia, also known as rabbit fever, is a disease caused by the bacterium Francisella tularensis. Tularemia is typically found in animals, especially rodents, rabbits, and hares. Tularemia is usually a rural disease and has been reported in all U.S. states except Hawaii.How do people become infected with tularemia?Typically, people become infected through the bite of infected insects (most commonly, ticks and deerflies), by handling infected sick or dead animals, by eating or drinking contaminated food or water, or by inhaling airborne bacteria.Created: 2009
-
This image depicts a Petri dish containing a buffered charcoal yeast extract (BCYE) agar medium, which had been inoculated with Gram-negative Francisella tularensis live vaccine strain (LVS) bacteria. F. tularensis is the pathogen responsible for causing the disease tularemia. This was the appearance of the colonial growth after 24 hours of incubation. What is tularemia?Tularemia, also known as rabbit fever, is a disease caused by the bacterium Francisella tularensis. Tularemia is typically found in animals, especially rodents, rabbits, and hares. Tularemia is usually a rural disease, and has been reported in all U.S. states except Hawaii.How do people become infected with tularemia?Typically, people become infected through the bite of infected insects (most commonly, ticks and deerflies), by handling infected sick or dead animals, by eating or drinking contaminated food or water, or by inhaling airborne bacteria.Created: 2009
-
Using methylene blue stain, this photomicrograph revealed the presence of Francisella tularensis bacteria, formerly known as Pasteurella tularensis. F. tularensis is the pathogen responsible for causing the disease tularemia.What is Tularemia?Tularemia is a potentially serious illness that occurs naturally in the United States. It is caused by the bacterium Francisella tularensis found in animals (especially rodents, rabbits, and hares).What are the Symptoms of Tularemia?Symptoms of tularemia could include: - sudden fever- chills - headaches- diarrhea - muscle - aches - joint pain - dry cough - progressive weaknessPeople can also catch pneumonia and develop chest pain, bloody sputum and can have trouble breathing and even sometimes stop breathing.Created: 1972
-
-
-
Thioploca is a filamentous sulfur-oxidizing bacterium, with the unusual characteristic of building sheaths around bundles of its filaments. They can travel up and down the sheaths by gliding motility. Thioploca can be found in sediments and also in microbial mats. These specimens are from Okinawa, Japan.
Thanks to the OIST Marine Science Support Section for hosting our visit in Nov 2017!
[taxonomy:genus=Thioploca]
-
Deposition authors: Minasov, G., Shuvalova, L., Dubrovska, I., Winsor, J., Scott, P., Anderson, W.F., Center for Structural Genomics of Infectious Diseases (CSGID);
Wikimedia Commons
Description: English: D-ribulose-5-phosphate 3-epimerase homododekamer + 24 SO4 (yellow-red) + 24 Cl (l.blue), Francisella tularensis. Date: 15 June 2016 (upload to Wikimedia Commons), 12 August 2009 (deposition at PDB). Source:
http://www.rcsb.org/pdb/explore/explore.do?structureId=3inp. Author: Deposition authors: Minasov, G., Shuvalova, L., Dubrovska, I., Winsor, J., Scott, P., Anderson, W.F., Center for Structural Genomics of Infectious Diseases (CSGID); visualization author:
User:Astrojan.
-
Description: English: chocolate agar medium with Francisella tularensis Polski: Agar czekoladowy z bakterią Francisella tularensis. Date: 2009. Source:
http://phil.cdc.gov/phil/details.asp. Author: CDC/ Megan Mathias and J. Todd Parker.
-
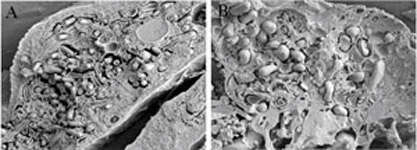
Summary.mw-parser-output table.commons-file-information-table,.mw-parser-output.fileinfotpl-type-information{border:1px solid #a2a9b1;background-color:#f8f9fa;padding:5px;font-size:95%;border-spacing:2px;box-sizing:border-box;margin:0;width:100%}.mw-parser-output table.commons-file-information-table>tbody>tr,.mw-parser-output.fileinfotpl-type-information>tbody>tr{vertical-align:top}.mw-parser-output table.commons-file-information-table>tbody>tr>td,.mw-parser-output table.commons-file-information-table>tbody>tr>th,.mw-parser-output.fileinfotpl-type-information>tbody>tr>td,.mw-parser-output.fileinfotpl-type-information>tbody>tr>th{padding:4px}.mw-parser-output.fileinfo-paramfield{background:#ccf;text-align:right;padding-right:0.4em;width:15%;font-weight:bold}.mw-parser-output.commons-file-information-table+table.commons-file-information-table,.mw-parser-output.commons-file-information-table+div.commons-file-information-table>table{border-top:0;padding-top:0;margin-top:-8px}@media only screen and (max-width:719px){.mw-parser-output table.commons-file-information-table,.mw-parser-output.commons-file-information-table.fileinfotpl-type-information{border-spacing:0;padding:0;word-break:break-word;width:100%!important}.mw-parser-output.commons-file-information-table>tbody,.mw-parser-output.fileinfotpl-type-information>tbody{display:block}.mw-parser-output.commons-file-information-table>tbody>tr>td,.mw-parser-output.commons-file-information-table>tbody>tr>th,.mw-parser-output.fileinfotpl-type-information>tbody>tr>td,.mw-parser-output.fileinfotpl-type-information>tbody>tr>th{padding:0.2em 0.4em;text-align:left;text-align:start}.mw-parser-output.commons-file-information-table>tbody>tr,.mw-parser-output.fileinfotpl-type-information>tbody>tr{display:flex;flex-direction:column}.mw-parser-output.commons-file-information-table+table.commons-file-information-table,.mw-parser-output.commons-file-information-table+div.commons-file-information-table>table{margin-top:-1px}.mw-parser-output.fileinfo-paramfield{box-sizing:border-box;flex:1 0 100%;width:100%}} Description: Coxiella burnetii-infected Vero cells (a) and Francisella tularensis-infected mouse macrophages (b) after chemical fixation, plunge freezing, fracturing, and viewing by cryo-SEM. Credit: NIAID. Date: 3 January 2013, 10:38. Source:
Freeze-Fractured Specimens. Author:
NIAID.
-
Description: English: Stained microphotograph of Thiomargarita namibiensis bacteria. Date: 2007. Source:
Commons - cropped and rotated. Author: NASA.
-
Description: Distribution of Thiomargarita namibiensis along the namibian coast. Date: 29 October 2007. Source: Own work, Namibia_sat.png. Author:
Denis Barthel. Other versions:
Image:Namibia_sat.png.
-
Description:
en:Francisella tularensis colonization on Cysteine Heart Agar after 72 hours. Obtained from the CDC
Public Health Image Library. Image credit: CDC/Larry Stauffer, Oregon State Public Health Laboratory (PHIL #1910), 2002. Source: This file is lacking source information. Please edit this file's description and provide a source. Author: This file is lacking author information.
-
Description: English: Beggiatoa-like filaments in underwater cave named "Grotta sulfurea", Capo Palinuro, Salerno (Italy). This mats is composed of filaments resembling in most part Beggiatoa, Thiothrix and Flexibacter. Date: 11 July 2008, 09:30:59. Source: Own work. Author:
Fabio Russo.
-
Description: English: Exended massive mats of some Beggiatoa-like filamentsin underwater cave named "Grotta sulfurea", Capo Palinuro, Salerno (Italy). This mats is composed of filaments resembling in most part Beggiatoa, Thiothrix and Flexibacter. Date: 11 July 2008, 09:30:31. Source: Own work. Author:
Fabio Russo.
-
Description: English: Some Beggiatoa-like filaments near a tiny hole in Capo Palinuro, Salerno, Italy. Date: 10 June 2007, 16:18:48. Source: Own work. Author:
Fabio Russo.
-
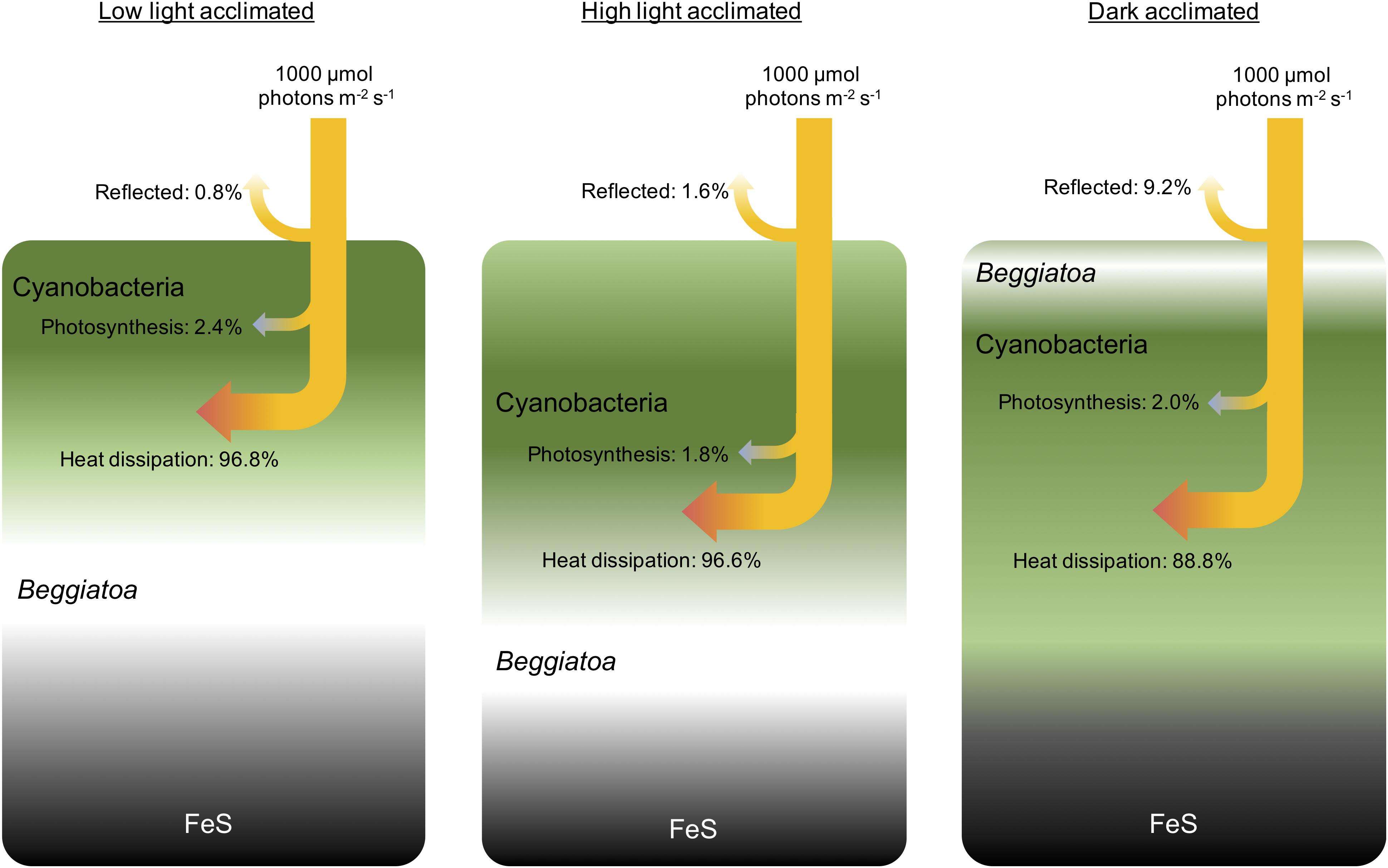
Description: English: Biomass distribution in a bacteria mat under different light conditions Conceptual drawing of the biomass distribution during different light acclimation scenarios and the corresponding radiative energy budgets of the entire photic zone measured under an incident photon irradiance of 1000 μmol photons m–2 s–1. The surface layers of the microbial mat were dominated by dense populations of motile cyanobacteria (Microcoleus sp. and other filamentous species) and filamentous sulfide oxidizing bacteria (Beggiatoa spp.). Beneath the narrow photic zone, a dark FeS-containing sediment layer was present. In low light, the upper layers became oxidized driving the O2-H2S interface and thus Beggiatoa deeper into the mat, while the motile cyanobacteria moved toward the mat surface under non-inhibitory light levels. In high light, a larger part of the biofilm is oxidized, Beggiatoa moved further down, while motile cyanobacteria in the top layers started to move downwards to avoid high photoinhibitory light levels at the microbial mat surface. In darkness, the entire mat became anoxic and the Beggiatoa moved to the mat surface forming a white layer on top of the cyanobacteria, to avoid the high levels of H2S and remain at the O2-H2S interface (Preisler et al., 2007), whereas the cyanobacteria remained in a dense layer underneath. Reference: Preisler, A., de Beer, D., Lichtschlag, A., Lavik, G., Boetius, A., and Jorgensen, B. B. (2007). "Biological and chemical sulfide oxidation in a Beggiatoa inhabited marine sediment". ISME J., 1: 341–353. Date: 14 May 2020, 03:21:21. Source:
[1] doi:
10.3389/fmars.2020.00359. Author: Mads Lichtenberg, Paulo Cartaxana and Michael Kühl.
-
Description: English: Drawing of Beggiatoa alba. Date: 10 December 2020, 18:04:20. Source: Own work. Author:
Fabio Russo.
-
Description: English: Little mat of Beggiatoa-like filaments associated to a source of thermal water rich in sulfur underwater, Massa Lubrense, Naples Italy. Date: 3 August 2012, 22:21:52. Source: Own work. Author:
Fabio Russo.